
Getting Started with because
Achaz von Hardenberg
2026-04-16
Source:vignettes/01_getting_started.Rmd
01_getting_started.RmdGetting started
because is an R package designed to easily perform causal inference with Bayesian Structural Equation Models in JAGS. The package integrates the methods proposed by von Hardenberg & Gonzalez-Voyer (2025) to fit Phylogenetic Bayesian Structural Equation Models (PhyBaSE) and extends them to other types of covariance structures (eg. spatial autocorrelation, genetic relatedness etc.).
because main features:
- Causal Inference with d-Separation: Testing conditional independencies implied by your causal model.
- Custom Priors and Mechanistic Constraints: Incorporating prior knowledge and mechanistic constraints.
-
Mediation Analysis: Decomposing
effects into direct and indirect components. Deterministic Nodes:
Interactions, Tresholds and mathematical transformations.
-
Phylogenetic Path Analysis: Using the Phylogenetic Bayesian
Structural Equation Model approach (PhyBaSE, von Hardenberg &
Gonzalez-Voyer, 2025)(through the
because.phybasemodule)
-
Phylogenetic Path Analysis: Using the Phylogenetic Bayesian
Structural Equation Model approach (PhyBaSE, von Hardenberg &
Gonzalez-Voyer, 2025)(through the
- Phylogenetic Missing Data Imputation*: Handling missing data using
Bayesian imputation (through the
because.phybasemodule).
- Phylogenetic Missing Data Imputation*: Handling missing data using
Bayesian imputation (through the
-
Alternative Covariance structures: Spatial, genetic,
social, or other correlation structures. Alternative dustribution
families: Modeling non-Gaussian data.
- Gaussian (continuous data)
- Binomial (binary/proportion data)
- Multinomial (unordered categorical data)
- Ordinal (ordered categorical data)
- Poisson (count data)
- Negative Binomial (overdispersed count data
- Zero inflated Poissson (ZIP) and negative binomial (ZINB)
- Advanced Model Specifications: random effects (mixed models, nested designs), polynomial terms, categorical predictors, measurement error, missing data.
- Hierarchical Data: Multi-level data with variables at different hierarchical levels.
- Latent Variables and MAG: Measurement error models and causal inference in the presence of latent variables using the MAG approach by Shipley & Douda (2021).
- Model Diagnostics: Checking convergence, comparing models, and interpreting results.
Quick Start
Installation
Before using because, you need to have JAGS (Just Another Gibbs Sampler) installed on your machine.
-
macOS:
brew install jagsor download from SourceForge. - Windows: Download installer from SourceForge.
-
Linux:
sudo apt-get install jags.
After installing JAGS, you can install because from
GitHub.
To install the stable release
(v1.2.6):
remotes::install_github("because-pkg/because@v1.2.6", build_vignettes = TRUE)To install the latest development version:
remotes::install_github("because-pkg/because", build_vignettes = TRUE)Finally, load the package:
Your First Model
The main function in because is because(), which
compiles and fits your specified Structural Equation Model in JAGS. The
function is very rich with functionalities allowing to model complex
models with different error structures and hierarchically structured
data. However, here, to show the basic workflow, we will use because()
to fit a simple linear model involving only two variables. Having only
two variables this is not a typical SEM being equivalent to a simple
linear regression which can not be used to infer causality, but it
serves to illustrate the basic usage of the package.
Let’s start simulating two variables X and Y where Y is correlated to X:
# set seed for reproducibility
set.seed(67)
# Simulate predictor
n <- 100
X <- rnorm(n = n, mean = 50, sd = 10)
# Generate response with the chosen intercept (alpha) and slope (beta)
alpha <- 20
beta <- 0.5
Y <- alpha + beta * X + rnorm(n, mean = 0, sd = 10)
# Combine into data frame
sim.dat <- data.frame(X, Y)
# Fit linear model with lm() function for comparison
summary(lm(Y ~ X, data = sim.dat))Now we can fit the same model using because. We need to specify the
structural equations (in this case only one) using R’s formula syntax
and then call because() passing the equation and the data
frame:
# Define the equations
equations <- list(Y ~ X)
# Fit the model
fit <- because(
equations = equations,
data = sim.dat
)
# View results
summary(fit)To check for convergence of the MCMC chains, you can look at the Rhat values in the summary output (should be < 1.1). You can also plot the trace plots of the MCMC samples:
# Plot trace plots
plot(fit$samples)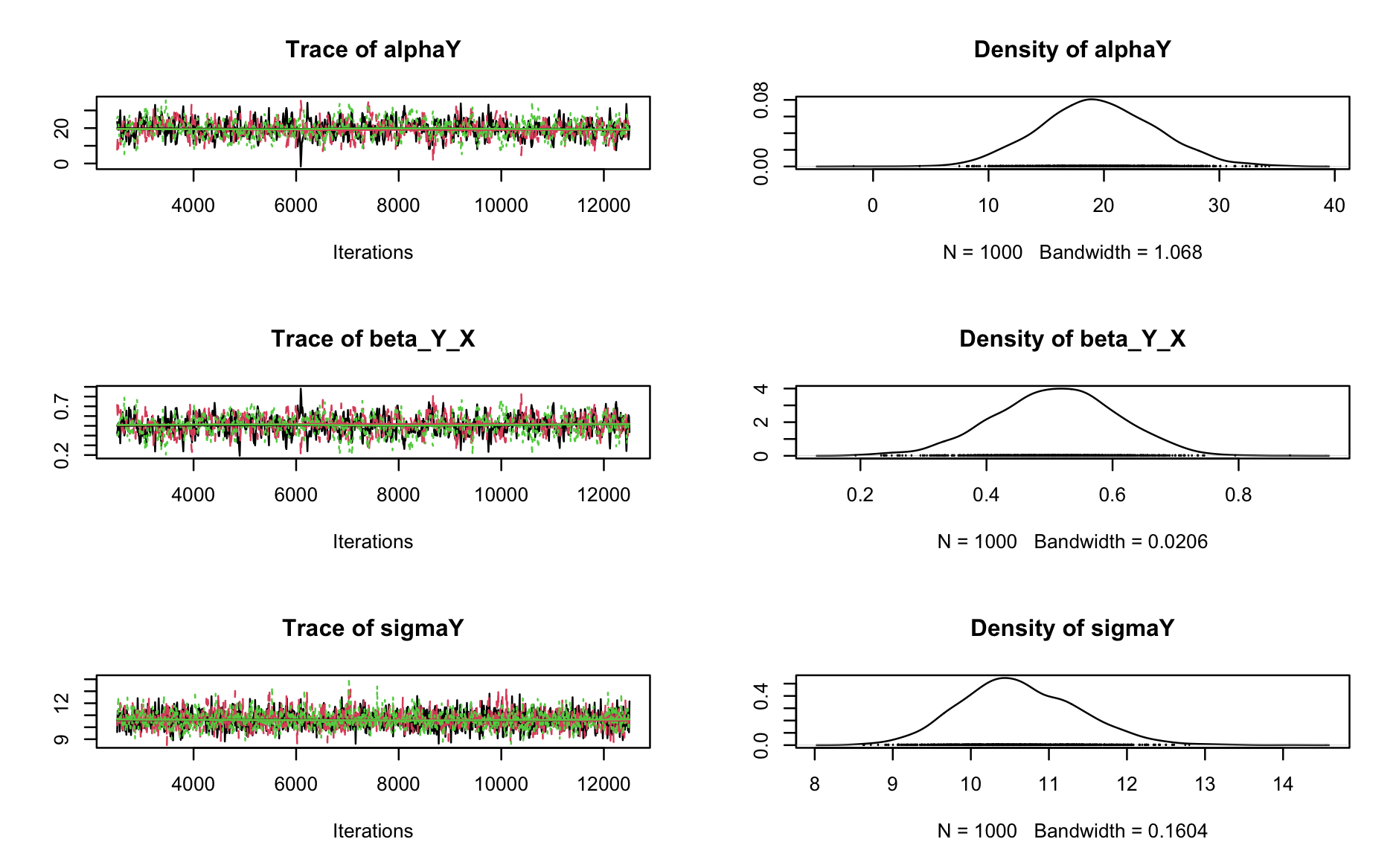
fit$samples is an MCMC object, so you can use all coda
functions to analyze and plot the MCMC samples.
You can see how because() translates your model into
JAGS syntax calling fit$model or, before fitting it, using
because_model():
# Generate JAGS model code
jags_model_code <- because_model(
equations = equations
)
cat(jags_model_code$model)When specifing the equations you can include multiple predictors as well as factors,interaction terms and polynomial terms following the conventional R formula syntax.
Next: Causal Inference with D-Separation
One of the main features of because is the ability to test your causal model’s fit to the data using d-separation tests. D-separation tests evaluate whether the conditional independencies implied by your causal model hold in the data (Shipley, 2016). If they do not, this suggests that your model may be misspecified and that you may need to add or remove paths. More details on d-separation tests, how to interpret them and a full tutorial can be found in the Causal inference with d-separation vignette.